Note
Go to the end to download the full example code.
Network assortativity and distance-preserving surrogates
This example demonstrates how to calculate network assortativity and generate distance-preserving surrogate connectomes for null hypothesis testing. We’ll use structural connectivity data and myelin maps from Bazinet et al. (2023) to explore how brain network topology relates to spatial patterns of cortical features:
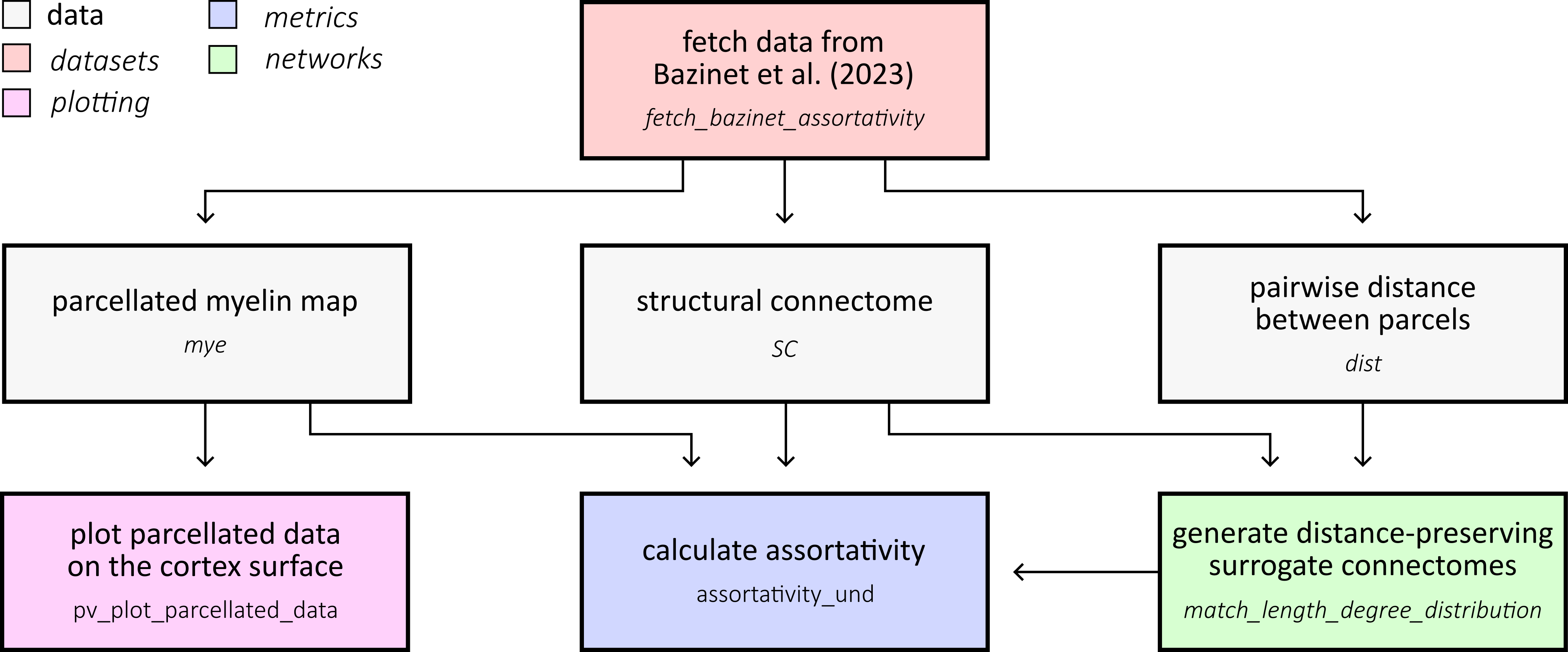
First, let’s fetch the structural connectivity and myelin data from Bazinet et al. (2023). This dataset contains a 400-parcel structural connectome with a corresponding distance matrix specifying the distance between parcel centroids. It also contains a T1w/T2w myelin map:
import numpy as np
import pickle
from netneurotools.datasets import fetch_bazinet_assortativity
bazinet_path = fetch_bazinet_assortativity()
with open(f'{bazinet_path}/data/human_SC_s400.pickle', 'rb') as file:
human_data = pickle.load(file)
SC = human_data['adj']
dist = human_data['dist']
myelin = human_data['t1t2']
print(f'Structural connectivity shape: {SC.shape}')
print(f'Distance matrix shape: {dist.shape}')
print(f'Myelin data shape: {myelin.shape}')
Please cite the following papers if you are using this function:
[primary]:
Vincent Bazinet, Justine Y Hansen, Reinder Vos de Wael, Boris C Bernhardt, Martijn P van den Heuvel, and Bratislav Misic. Assortative mixing in micro-architecturally annotated brain connectomes. Nature Communications, 14(1):2850, 2023.
[fetch_single_file] Downloading data from
https://github.com/netneurolab/bazinet_assortativity/archive/baab96d173b8c5a2656
67a4259cf228713dabcc7.tar.gz ...
[fetch_single_file] Downloaded 14155776 of ? bytes.
[fetch_single_file] Downloaded 27795456 of ? bytes.
[fetch_single_file] Downloaded 41426944 of ? bytes.
[fetch_single_file] Downloaded 55320576 of ? bytes.
[fetch_single_file] Downloaded 59252736 of ? bytes.
[fetch_single_file] Downloaded 63184896 of ? bytes.
[fetch_single_file] Downloaded 67117056 of ? bytes.
[fetch_single_file] Downloaded 75030528 of ? bytes.
[fetch_single_file] Downloaded 94642176 of ? bytes.
[fetch_single_file] Downloaded 105914368 of ? bytes.
[fetch_single_file] Downloaded 108797952 of ? bytes.
[fetch_single_file] Downloaded 111943680 of ? bytes.
[fetch_single_file] Downloaded 114827264 of ? bytes.
[fetch_single_file] Downloaded 118759424 of ? bytes.
[fetch_single_file] Downloaded 133439488 of ? bytes.
[fetch_single_file] Downloaded 136323072 of ? bytes.
[fetch_single_file] Downloaded 139206656 of ? bytes.
[fetch_single_file] Downloaded 142876672 of ? bytes.
[fetch_single_file] Downloaded 147857408 of ? bytes.
[fetch_single_file] Downloaded 150740992 of ? bytes.
[fetch_single_file] Downloaded 155713536 of ? bytes.
[fetch_single_file] Downloaded 159129600 of ? bytes.
[fetch_single_file] Downloaded 163586048 of ? bytes.
[fetch_single_file] Downloaded 168566784 of ? bytes.
[fetch_single_file] Downloaded 171704320 of ? bytes.
[fetch_single_file] Downloaded 174850048 of ? bytes.
[fetch_single_file] Downloaded 177995776 of ? bytes.
[fetch_single_file] Downloaded 195035136 of ? bytes.
[fetch_single_file] Downloaded 204480512 of ? bytes.
[fetch_single_file] Downloaded 207355904 of ? bytes.
[fetch_single_file] Downloaded 210239488 of ? bytes.
[fetch_single_file] Downloaded 213917696 of ? bytes.
[fetch_single_file] Downloaded 218112000 of ? bytes.
[fetch_single_file] Downloaded 220995584 of ? bytes.
[fetch_single_file] Downloaded 228065280 of ? bytes.
[fetch_single_file] Downloaded 230957056 of ? bytes.
[fetch_single_file] Downloaded 233840640 of ? bytes.
[fetch_single_file] Downloaded 236986368 of ? bytes.
[fetch_single_file] Downloaded 241180672 of ? bytes.
[fetch_single_file] Downloaded 244326400 of ? bytes.
[fetch_single_file] Downloaded 252444672 of ? bytes.
[fetch_single_file] Downloaded 278601728 of ? bytes.
[fetch_single_file] Downloaded 305405952 of ? bytes.
[fetch_single_file] Downloaded 327163904 of ? bytes.
[fetch_single_file] Downloaded 348389376 of ? bytes.
[fetch_single_file] Downloaded 369369088 of ? bytes.
[fetch_single_file] Downloaded 390602752 of ? bytes.
[fetch_single_file] Downloaded 412098560 of ? bytes.
[fetch_single_file] Downloaded 433332224 of ? bytes.
[fetch_single_file] Downloaded 454303744 of ? bytes.
[fetch_single_file] Downloaded 476585984 of ? bytes.
[fetch_single_file] Downloaded 499130368 of ? bytes.
[fetch_single_file] Downloaded 519577600 of ? bytes.
[fetch_single_file] Downloaded 541859840 of ? bytes.
[fetch_single_file] Downloaded 562561024 of ? bytes.
[fetch_single_file] Downloaded 585637888 of ? bytes.
[fetch_single_file] ...done. (59 seconds, 0 min)
Downloaded ds-bazinet_assortativity to /home/runner/nnt-data
/home/runner/work/netneurotools/netneurotools/examples/plot_assortativity.py:34: DeprecationWarning: numpy.core.numeric is deprecated and has been renamed to numpy._core.numeric. The numpy._core namespace contains private NumPy internals and its use is discouraged, as NumPy internals can change without warning in any release. In practice, most real-world usage of numpy.core is to access functionality in the public NumPy API. If that is the case, use the public NumPy API. If not, you are using NumPy internals. If you would still like to access an internal attribute, use numpy._core.numeric._frombuffer.
human_data = pickle.load(file)
Structural connectivity shape: (400, 400)
Distance matrix shape: (400, 400)
Myelin data shape: (400,)
Let’s visualize the structural connectivity matrix. This shows the weighted connections between all pairs of brain regions:
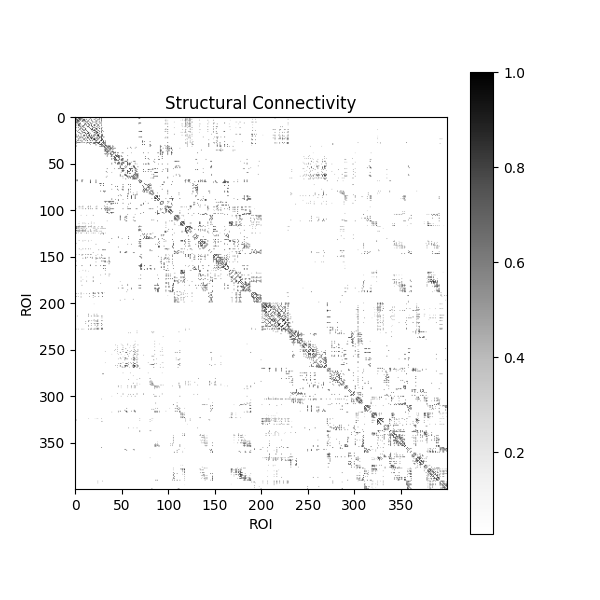
<matplotlib.colorbar.Colorbar object at 0x7f948643a810>
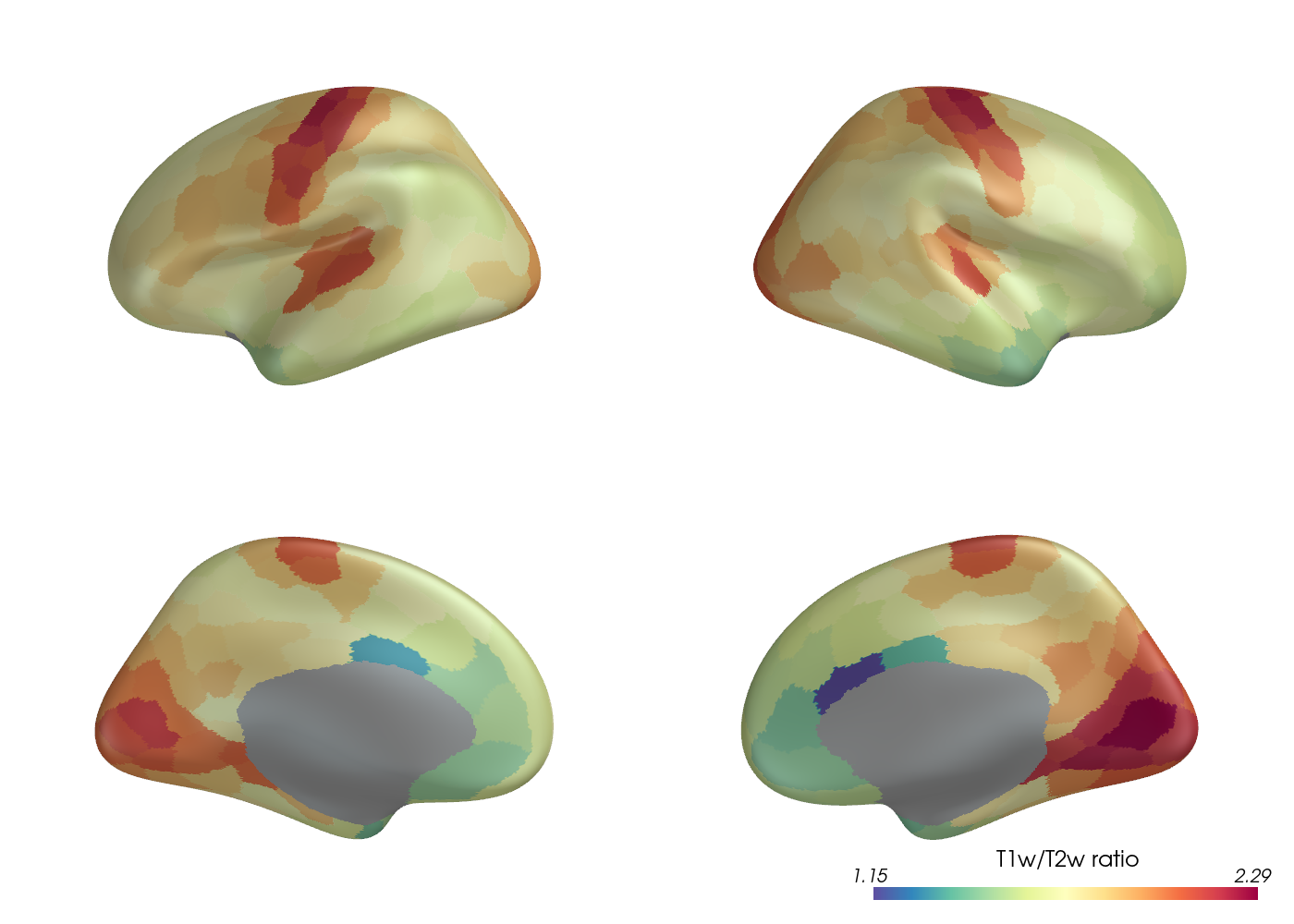
Let’s visualize the parcellated myelin data on the cortical surface. The structural connectivity and myelin data have been parcellated with the Schaefer (400 nodes) parcellation.
from netneurotools.plotting import pv_plot_parcellated_data
# This is necessary for headless rendering when building the sphinx gallery.
# If you are running this code locally, you might not need this line.
import os
os.environ["VTK_DEFAULT_OPENGL_WINDOW"] = "vtkOSOpenGLRenderWindow"
pv_plot_parcellated_data(myelin,
'schaefer400x7',
cmap='Spectral_r',
clim=[np.min(myelin), np.max(myelin)],
cbar_title='T1w/T2w ratio',
lighting_style='plastic',
jupyter_backend="static")
Please cite the following papers if you are using this function:
[primary]:
Alexander Schaefer, Ru Kong, Evan M Gordon, Timothy O Laumann, Xi-Nian Zuo, Avram J Holmes, Simon B Eickhoff, and BT Thomas Yeo. Local-global parcellation of the human cerebral cortex from intrinsic functional connectivity mri. Cerebral cortex, 28(9):3095–3114, 2018.
[fetch_single_file] Downloading data from
https://files.osf.io/v1/resources/udpv8/providers/osfstorage/673273c54b79ea9a506
2f4b3 ...
[fetch_single_file] ...done. (2 seconds, 0 min)
Downloaded atl-schaefer2018 to /home/runner/nnt-data
<PIL.Image.Image image mode=RGB size=1400x1000 at 0x7F9486439DC0>
<pyvista.plotting.plotter.Plotter object at 0x7f9485b36330>
Network assortativity measures the tendency for nodes with similar features to be connected. Let’s calculate the assortativity of the myelin distribution with respect to the structural connectivity network:
from netneurotools.metrics import assortativity_und
assort = assortativity_und(myelin, SC, use_numba=False)
print(f'Network assortativity: {assort:.3f}')
Network assortativity: 0.463
We can visualize this by creating a scatterplot showing the myelin values of connected node pairs, with point colors indicating connection strength:
SC_E = SC > 0
Xi = np.repeat(myelin[np.newaxis, :], 400, axis=0)
Xj = np.repeat(myelin[:, np.newaxis], 400, axis=1)
sort_ids = np.argsort(SC[SC_E])
fig, ax = plt.subplots(figsize=(6, 6))
scatter = ax.scatter(
Xi[SC_E][sort_ids],
Xj[SC_E][sort_ids],
c=SC[SC_E][sort_ids],
cmap='Greys',
s=3,
rasterized=True
)
ax.set(xlabel='Myelin (node i)', ylabel='Myelin (node j)')
ax.set_title(f'Assortativity = {assort:.3f}')
fig.colorbar(scatter, ax=ax, label='Connection strength')
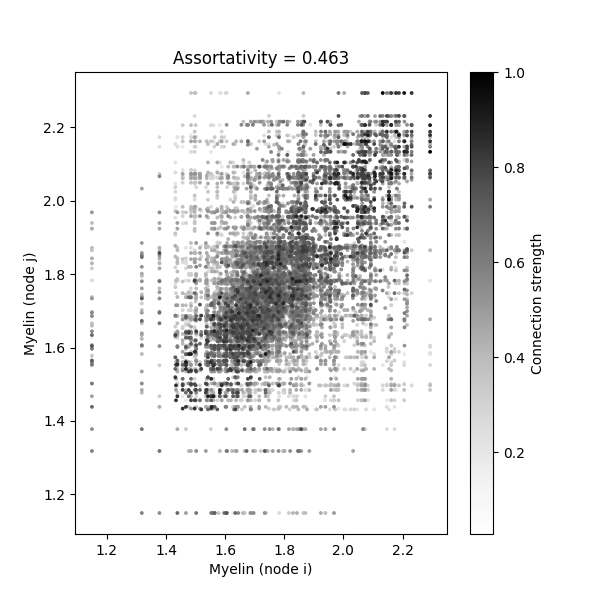
<matplotlib.colorbar.Colorbar object at 0x7f9484137020>
To test whether this assortativity is significant, we need null models that preserve key topological properties. We’ll generate distance-preserving surrogate networks that maintain the degree distribution and spatial embedding of connections. For this, we need the empirical structural connectome and a matrix specifying the distance between each parcel.
Note: Generating many surrogates can be time-consuming. For this example, we’ll create just a few surrogates:
from netneurotools.networks import match_length_degree_distribution
n_surrogates = 10
surr_all = np.zeros((n_surrogates, 400, 400))
for i in range(n_surrogates):
surr_all[i] = match_length_degree_distribution(SC, dist)[1]
print(f'Generated {n_surrogates} surrogate networks')
Generated 10 surrogate networks
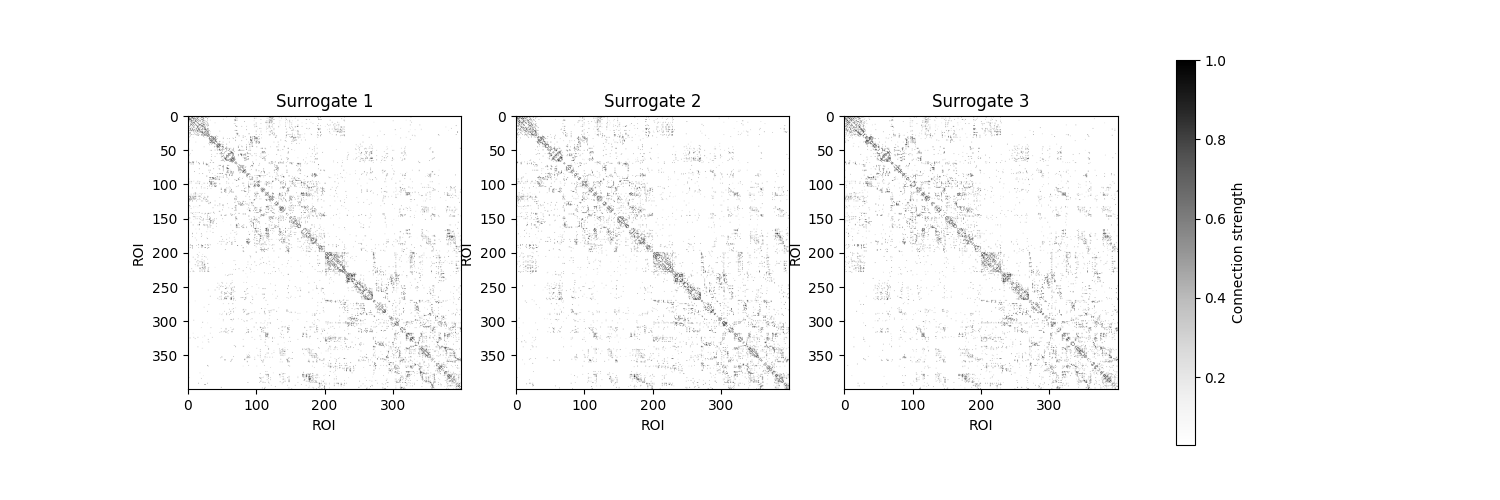
Let’s visualize a few of these surrogate connectomes to see how they compare to the empirical network:
fig, axes = plt.subplots(1, 3, figsize=(15, 5))
for i, ax in enumerate(axes):
im = ax.imshow(
surr_all[i],
cmap='Greys',
vmin=surr_all[i][surr_all[i] > 0].min(),
vmax=surr_all[i].max()
)
ax.set_title(f'Surrogate {i + 1}')
ax.set(xlabel='ROI', ylabel='ROI')
fig.colorbar(im, ax=axes.ravel().tolist(), label='Connection strength')
<matplotlib.colorbar.Colorbar object at 0x7f94841d4fe0>
Now we’ll calculate the assortativity for each surrogate network to create a null distribution:
assort_surr = np.zeros(n_surrogates)
for i in range(n_surrogates):
assort_surr[i] = assortativity_und(myelin, surr_all[i])
print(f'Mean surrogate assortativity: {np.mean(assort_surr):.3f}')
print(f'Std surrogate assortativity: {np.std(assort_surr):.3f}')
Mean surrogate assortativity: 0.400
Std surrogate assortativity: 0.006
Finally, we can compare the empirical assortativity to the null distribution using a boxplot. This visualization shows whether the observed assortativity is significantly different from what we’d expect by chance:
fig, ax = plt.subplots(figsize=(4, 6))
flierprops = dict(
marker='+',
markerfacecolor='lightgray',
markeredgecolor='lightgray'
)
bplot = ax.boxplot(
assort_surr,
widths=0.3,
patch_artist=True,
showfliers=True,
showcaps=False,
flierprops=flierprops
)
for box in bplot['boxes']:
box.set(facecolor='lightgray', edgecolor='black')
for median in bplot['medians']:
median.set(color='black', linewidth=2)
ax.scatter(1, assort, color='red', s=100, zorder=3, label='Empirical')
ax.set_xticks([1])
ax.set_xticklabels(['Surrogates'])
ax.set_ylabel('Assortativity')
ax.set_title('Empirical vs. Null Distribution')
ax.legend()
ax.set_xlim(0.7, 1.3)
plt.tight_layout()
The red dot shows the empirical assortativity, while the boxplot shows the distribution of assortativity values from the distance-preserving surrogate networks. If the empirical value falls outside the null distribution, this suggests that the observed pattern of myelin assortativity is not explained by the spatial constraints and topology of the connectome alone.
The limitation is equally important: geometry-aware nulls still depend on the quality of the upstream surface / centroid information, and generating large surrogate ensembles can be computationally expensive.
Total running time of the script: (1 minutes 15.615 seconds)